Chapter 9: Variofit


The variofit application is used to fit model semivariograms to experimental and
jackknifed experimental semivariograms (Generated by vario). This can be done
manually or automatically using least-squares regression or latin-hypercube sampling
techniques. Ergodic variations of the model semivariogram from simulation series may
also be evaluated.
The variofit application is composed of three sections; the main menu-bar, the
status and log text area, and the drawing or graph area. The menu-bar is used to select
all vario commands, the log/status area is used by the program to report important
messages or results, and the drawing area is the display area for the semivariogram
graph.
Menu Items
Examples
Command Line Arguments
File Formats
Mathematics
Bibliography
The main menu controls nearly all the program operations; files can be opened
and saved, graphics can be plotted, the appearance of the graphic can be modified, help
can be requested, and the results can be sent to the printer. For variofit there are
thirteen items on the main menu:
File,
Regression,
Jackknife,
Latin-Hyperrcube,
Series,
Graph,
Log,
Plot, and
Help,
(Figure 9.1). File controls file handling (opening, saving, naming files), directs printing, and allows the user to quit the application. Regression specifies the parameters for manually or automatically fitting a model semivariogram to an experimental semivariogram. Jackknife specifies some of the parameters pertinent to viewing jackknifed semivariograms. Latin-Hypercube specifies the parameters for
fitting multiple model semivariograms to a jackknifed experimental semivariograms.
Series loads a simulation series of experimental semivariograms (calculated in vario
(Chapter 8), and defines relevant parameters for viewing the data. Graph is used to
define details about the graph border, fonts, label, mesh, and line styles. Log allows the
user to save the text messages and results printed to the log/status window. Plot plots
the graph. Help gives the user a selection of pop-up help topics. Each menu item is
fully described below with all the available options.
 Figure 9.1
Figure 9.1
The File sub-menu options control file and print handling, and exiting the
program. The options include Open, View Data, View Results, Save, Save as, Save
Preferences, Print Setup, Print, Quit, and Quit Without Saving.
Selecting File:Open generates a pop-up dialog which allows the user to select an
existing data file. This dialog operates exactly as the Open:File dialog in Chapter 5
(plotgraph Figure 5.2). However, unlike plotgraph the default data file extension is
*.gam (See vario, Chapter 8, for data file format).
File:View Data pops up a simple screen editor showing the file that the current
experimental semivariogram or jackknifed experimental semivariogram is based upon.
File:View Results pops up a simple screen editor showing the file containing the
last saved results of a model fit calculation. See Output Data File Format Section for
data file format.
File:Save saves the results of the latest calculation to a file. If a save file has
already been opened, the data are simply saved. If a save file has not been selected yet,
a pop-up dialog similar to that used in File:Open (Figure 5.2) is created. The main
difference between the Open and the Save dialog is that to save a file, the file does not
have to pre-exist. For a description of how the dialog works, see the Open section above
and substitute Save for Open wherever appropriate.
The graph lines will be saved using one of the two file formats specified in the
"File Output" section (single format, and multiple jackknifed format).
File:Save as is identical to File:Save described above, when a file has not been
selected yet. This option can be used to save the file for the first time, or save results to
a new file.
When using programs with many user options, it is not possible for the program
to always pick reasonable default values for each parameter or input variable. For this
reason preference files were created (See Appendix C). These allow the user to define a
unique set of "defaults" applicable to the particular project. When File:Save Preferences
is selected, variofit determines how all the input variables are currently defined and
writes them to the file "variofit.prf."
- WARNING: if "variofit.prf" already exists, you will be warned that it is about to be
over-written. If you do not want the old version destroyed you must move it to a
new file (e.g. the UNIX command mv variofit.prf variofit.old.prf would be
sufficient). When you press OK the old version will be over-written! Moving the
old file cannot be done currently from within the application. To rename the you
will have to execute the UNIX mv command from a UNIX prompt in another
window. If "variofit.prf" does not exist in the current directory, it is created.
This is an ASCII file and can be edited by the user. See Appendix C for details.
File:Print Setup works exactly as explained in Chapter 5.
File:Print generates a Postscript file of the calculated spatial measure, and
depending on how the print options are define in Print Setup, directs this file to the
specified print queue, or to the specified file.
File:Quit terminates the program, but if additions have been made to the graph,
the user will first be queried to supply a file to save the changes in.
File:Quit Without Saving terminates the program regardless of any additions to
the graph. Once pressed there is no option to change your mind.
[TOP] [SYNTAX]
The Regression menu option is used to fit a single model semivariogram to an
experimental semivariogram. Manual an automatic fitting routines are available. For a
description of the mathematics used for defining or for automatically calculating a
model semivariogram, see the Mathematics Section below.
There are two options for model calculation, Automatic and Manual. The
Automatic method a least-squares regression method and is useful for getting an
approximate model fit. For many solutions this automated fit may give reasonable
results, but rarely is the automated least-squares fit the best fit for the model
semivariogram. This method usually yields satisfactory results, but the algorithm still
needs work, and sometimes it gives very poor results. Fitting a model semivariogram is
a bit of an art, and a geostatistician may prefer a slightly different model. The
automated solution is, though, good for making a first approximation. The Manual
solution method is used when the user wishes to improve upon the automatic fit, or
define the model directly. Here the range, sill, and nugget values can be adjusted as
desired, and the model semivariogram can be displayed with the experimental
semivariogram.
Automatic:
To automatically fit a model semivariogram to an experimental semivariogram,
several steps have to be followed:
- 1) Open a experimental semivariogram data file (File:Open).
2) Select Regression:Calculation Method:Automatic
3) Display the Regression:Parameters dialog.
4) Select the desired Solution Method.
5) Select the number of model nests desired and the model type for each nest.
6) Set the Starting Range and the Ending Range for the iterative solutions to iterate
upon.
7) Set the number of iterative solutions (Calculation Steps - Division) for the
algorithm to iterate upon.
8) Select Regression:Calculate from the main menu-bar to start the regression.
While the regression is being calculated, the range increment for nests 4, 3, and 2 will
be displayed in the status log window along with the largest error calculated for all
increments of the first nest range. Once the regression is complete, the "best-fit" model
semivariogram will be drawn in the graphics area and the optimum range, sill, and
nugget values will be updated within the pop-up regression dialog.
- NOTE: Any user input values for Range, Sill, or Nugget will be overwritten when an
automatic solution is calculated.
WARNING: There are still bugs in this algorithm and the optimized solution may be
an extremely poor fit!
WARNING: The automatic solver uses an iterative approach (there are more
unknowns than equations) to optimize the model solution. Because of this it is
not recommended that this solver be used to fit three or four nested structures.
Using the default value for Division Steps (100) a two-nested model takes
approximately 100 times as long to solve as a single-nested model (i.e. 30 sec.
vs. 0.3 sec). For the same data set, a three-nested model would take 50
minutes, and a four-nested model several days!
Manual:
To manually fit a model semivariogram, two options are available. Parameter
values may be entered directly, or a fitting tool with slider bars can be used (discussed
below). To fit a model by directly entering values, follow these steps:
-
1) Open a experimental semivariogram data file (File:Open).
2) Select Regression:Calculation Method:Manual.
3) Display the Regression:Parameters dialog.
4) Select the number of model nests desired and the model type for each nest.
5) Define the desired values for each appropriate Range, Sill, and Nugget value.
6) Press the Apply button to enter all values in the dialog.
7) Select Regression:Calculate from the main menu-bar to start the regression.
The described model semivariogram will be displayed in the graphics area,
superimposed over the experimental semivariogram (Figures 9.2 and 8.3). To refine the
model fit further, repeat steps 4) through 7).
 Figure 9.2
Figure 9.2
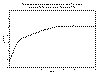 Figure 9.3
Figure 9.3
To fit a model manually using the slider fitting tool, select the Manual
Calculation Method and select the Regression:Slider Fitting menu item. This will
produce the dialog shown in Figure 9.4.
 Figure 9.4
Figure 9.4
Selecting Regression:Parameters generates the pop-up dialog shown in Figure 9.5. This dialog describes everything necessary to fit a model semivariogram to an
experimental semivariogram. Up to four nested structures can be modeled. For each nested structure (at least one is required) the Range and Sill values may be entered,
and the model type defined (Linear, Linear with Sill, Spherical, Gaussian, Exponential,
and Logarithmic are valid model types). No Model indicates that the defined model nest
and any higher nest will not be evaluated (The first column on the left defines the first
nest structure, the fourth column on the right defines the fourth nest structure). The
Nugget may also be defined.
 Figure 9.5
Figure 9.5
Because the graph can get very cluttered when there are several experimental
semivariograms displayed, the Visible Models button was added. When pressed, the
pop-up dialog shown in Figure 9.6 is created. With this dialog, different experimental
semivariograms can be turned on and off.
 Figure 9.6
Figure 9.6
When an automatic solution is desired several other parameters need to be
defined. The automated method uses an iterative solver because there are more
unknown's than equations. Because of this, some limits must be placed on the
iteration. By default, the maximum possible range of the data set (Starting Range) is
considered to be the maximum diagonal distance across the data set, and the minimum
possible range is considered 0.0 (Ending Range). In the iterative solution, this possible
range of Range values is divided into 100 (default value for Calculation Steps - Divisions)
subdivisions. For the first iteration, a "best-fit" model will be calculated assuming the
maximum range. For the next iteration, the range will be set to the maximum range -
(maximum range / 100.0). This will be repeated until the minimum range (0.0) is used.
For each iteration, the least-squares error for each model semivariogram will be saved.
The model semivariogram with the smallest squared-error, is considered the best-fit. In
many situations, it may be reasonable to modify these values. For example, by
examining the experimental semivariogram in Figure 9.2, it is clear the range is less
than 65 meters. If the data sets maximum possible range was 500 meters a great deal
of computational effort would be wasted examining ranges greater than 70 meters. A
similar argument can be made for the Ending Range. The number of Divisions defines
the fineness of the steps used to determine the range. If the range is fairly narrow fewer
divisions may be reasonable; if a very wide range is possible more divisions may be
necessary.
-
NOTE: The program supports only models with up to four structures. It is felt that
only in an extremely unusual case would even four structures be justified, much
less five or more.
NOTE: The range for each model structure must be between the Starting and
Ending Range parameters. Do not over constrain the solution.
NOTE: For multi-nested structures, the number of Calculation Steps- Divisions is
the main control on how long the automatic solution takes. Reducing the
number of divisions can reduce the solution time significantly!
The Slider Fitting tool shown the Figure 9.4 can be used in the Manual
calculation mode. Using this tool, the Nugget, Cnestnest values can be fit to
the experimental semivariogram. When the slider bars are moved the current
parameter value will be used to a model semivariogram on the graphics display. By
moving the slider bars (with a little practice), and selecting the appropriate nest, the model semivariogram can be moved and adjusted to overlay the experimental
semivariogram.
When an automatic solution is selected, a least-squared method must be
defined. This method defines how the least-squares regression algorithm will weight
and consider errors between the fit model and the experimental semivariogram. The
options are displayed in a pop-up dialog (Figure 9.7) and explained below:
 Figure 9.7
Figure 9.7
- Least-Squares - This method fits a "best-fit" model curve to the data. No
adjustments are made for the nugget, variance, or the relative number of pairs
per lag.
Nugget LS - This method determines the nugget by determining the y-intercept of a
line drawn through the first two 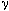 (h) lag values. Once the nugget is determined
the standard least-squares approach used above is applied.
(h) lag values. Once the nugget is determined
the standard least-squares approach used above is applied.
Variance LS - This method defines the sill as equaling the variance of the data set.
This forces the model curve to honor the data set variance; this is also
commonly done by geostatisticians when manually fitting model
semivariograms. Once the nugget is determined the standard least-squares
approach used above is applied.
Nugget & Variance LS - This method combines the Nugget LS and Variance LS
methods described above.
Weighted LS - This method is a normal least-squares regression, except that it
weights the error for each lag, based on the number of sample pairs used to
generate that lag 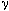 (h) value (i.e., this method presumes that it is more important
to honor a data point representing 500 data pairs than one representing 3 data
pairs). The previous four methods do not consider the number of data pairs,
therefore they weight all lag values equally.
(h) value (i.e., this method presumes that it is more important
to honor a data point representing 500 data pairs than one representing 3 data
pairs). The previous four methods do not consider the number of data pairs,
therefore they weight all lag values equally.
Selecting the Regression:Calculate menu option will cause the program to
calculate and plot a single model semivariogram based on how the parameters are set
in the Regression:Modify dialog (Figure 9.5). When the calculation is complete the
model parameters and Mean Square Error (MSE) are printed to the log/status window.
A word of caution; if nested structures are being calculated, particularly three or four,
this procedure may take a very long time if the Automatic calculation method is used.
Before you use this may nests, examine the experimental semivariogram carefully;
rarely are this many nested structures justified.
-
NOTE: This Regression is not equivalent to the regression option described in
Chapter 5 for plotgraph.
[TOP] [SYNTAX]
When a series of jackknifed semivariogram models is loaded (See vario, Chapter 8), various information can be displayed about the group of experimental
semivariograms.
Jackknife:Modify creates the pop-up dialog shown in Figure 9.8. This dialog
allows the user to specify what information is displayed about the jackknifed
experimental semivariograms. Draw Intermediate Results determines whether the
experimental semivariograms based on the number of data points minus one are
displayed. Draw Unjackknifed Solution determines whether the experimental
semivariogram based on all of the data points is displayed. Draw Error-Bars determines
whether the vertical error-bars representing the confidence limits of the  (h) value for
each lag are drawn. Draw Error-Bands determines whether the horizontal error-bars
representing the confidence limits of the lag distance are drawn. Draw Range of
Jackknifed Variance's: because each jackknifed experimental semivariogram is based
on a slightly different set of data points the variance for each solution is also different;
this option determines whether this variation is drawn. Finally, the confidence level for
the error-bars can be specified; 95% is the default.
(h) value for
each lag are drawn. Draw Error-Bands determines whether the horizontal error-bars
representing the confidence limits of the lag distance are drawn. Draw Range of
Jackknifed Variance's: because each jackknifed experimental semivariogram is based
on a slightly different set of data points the variance for each solution is also different;
this option determines whether this variation is drawn. Finally, the confidence level for
the error-bars can be specified; 95% is the default.
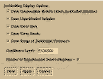 Figure 9.8
Figure 9.8
Jackknife:Individual Display creates the dialog shown in Figure 9.9 and allows
the user to display individually each jackknifed experimental semivariogram Figure
8.10. When the dialog is destroyed, all of the jackknifed semivariograms and other
relevant information is redisplayed. The jackknifed experimental semivariograms can
be displayed by number ID (ID number set by order of occurrence in data file), or they
can be stepped through by pressing the Next and Previous buttons.
 Figure 9.9
Figure 9.9
 Figure 9.10
Figure 9.10
[TOP] [SYNTAX]
Latin-Hypercube sampling is used in this package to select a limited number,
but a representative group of model semivariograms which honor the confidence limits
of a series of jackknifed experimental semivariograms (See Mathematics Section for
theory behind this approach). This approach is not meant for use on single
experimental semivariograms, or best estimate kriging. The model semivariogram
output from this technique is used for input into the SISIM3D stochastic conditional
simulation program (Chapter 14 ).
Latin-Hypercube:Modify generates the pop-up dialog shown in Figure 9.11. This
dialog is used to specify the parameters which control the latin-hypercube sampling
process.
 Figure 9.11
Figure 9.11
Unlike the Regression:Parameters dialog however, only two nested structures
can be modeled. Because of the lack of data inherent when this process is used,
modeling even two structures is probably suspect. For most cases there is probably
little justification for using more than one structure. As with the Regression:Parameters
dialog No Model, Linear, Linear with Sill, Spherical, Exponential, Gaussian, and
Logarithmic models can be used for each structure. Instead of specifying single Range,
Sill, and Nugget values though, the range of values must be specified, because this is a
randomized model fitting procedure.
For each variable; Range, Sill and Nugget, a Minimum and Maximum value must
be specified (The Minimum may equal the Maximum range, e.g. a constant is implied).
This specifies the range of allowable or reasonable values. These entries are preset with
default values:
-
Range: The Minimum Range is set by the longest lag that has its confidence interval
fall completely below the confidence interval of the jackknifed experimental
semivariogram variances (No lower lag values can fall above the variance either),
or failing this, to the minimum confidence interval range of the shortest lag
(Realistically, it might be best to set this value to 0.0 in this case since the
possibility of pure nugget effect cannot be disregarded) (Figure 9.12). The
Maximum Range is set by the shortest lag that has it confidence interval that lies
completely above the variance confidence interval. In many models this never
occurs, and the Maximum Range is set to the maximum extent of the maximum
lag confidence interval (Figure 9.12, the Maximum Range is set to 436 feet).
 Figure 9.12
Figure 9.12
Variance: The Minimum Variance is set to the lower bound of the jackknifed
experimental semivariogram confidence interval, and the Maximum Variance is
set to the upper bound (Figure 9.12, Minimum = 0.2613, Maximum = 0.2841).
Nugget: The Minimum Nugget is set to 0.0. The Maximum Nugget is set to the upper
bound of the shortest lag confidence interval, or the upper bound of the
jackknifed experimental semivariogram confidence interval; which ever is the
least (Figure 9.12, Minimum = 0.0, Maximum = 0.2762).
If the Minimum and Maximum entries appear inappropriate, the user is free to alter their
values. There are many justifications for doing so.
Each interval (Range, Variance, and Nugget) can be divided into one to ten
Divisions; by default these Divisions are of equal width. If non-equal Division intervals
are desired, press the Define button for the appropriate parameter. When this is done,
one of the pop-up dialogs shown in
Figures 9.13a,
8.13b, and
8.13c
will be created. In
this dialog the desired absolute division intervals can be assigned. If the Range
Increments have been adjusted and equal increments are desired, press the Set
Divisions Equal button. By default the Range and Nugget are divided into four Divisions
and the Variance one division. In this case, when the Latin-Hypercube sampled
semivariograms are calculated (Select the Latin-Hypercube:Calculate main menu-bar
option), 16 model semivariograms will be calculated (4 Range x 1 Variance x 4 Nugget
divisions x Number of Solutions per Division). An example is shown in Figure 9.14a. For
each model semivariogram, the model Range, Variance, and Nugget is randomly selected
from within each Range Interval (Equally Spaced selections from within the Range
Interval are Possible in addition to the Random Range Selection Option).
 Figure 9.13a,
Figure 9.13a,
 Figure 9.13b, and
Figure 9.13b, and
 Figure 9.13c
Figure 9.13c
 Figure 9.14a
Figure 9.14a
- NOTE: As long as no parameters are changed in this dialog, the randomly selected
model semivariograms will always be exactly the same; this is because the same
initial Randomize Seed is used. This allows the user to repeat previous work.
To generate a new series of random model semivariograms, select a new
Randomize Seed (any integer value).
Just because 16 model semivariograms have been calculated, this does not mean that
they honor the confidence intervals of the jackknifed experimental semivariogram. As
shown in Figure 9.14a three model semivariograms (dashed lines) do not honor the
confidence intervals of the shortest lag! These models honor the parameter values specified (Range, Variance, and Nugget), but they do not honor the jackknifed
experimental semivariogram and should be disregarded.
- NOTE: Apparently invalid model semivariograms are drawn, because in some cases
the models are reasonable, but the decision algorithm in the software is overly
sensitive. Carefully examine each model before discarding it.
Several options allow the software to check for, and eliminate invalid models. Note, if a
model is eliminated a new model is not calculated to replace it. If the Check Solutions
Within Range Area is active, the software will attempt to determine which, if any models
do not honor the experimental data. If Reject Invalid Solutions is also selected, invalid
models will not be drawn (Figure 9.14b), otherwise they will be drawn with dashed lines (Figure 9.14a).
 Figure 9.14b
Figure 9.14b
- NOTE: Draw Valid Range Area is not currently installed.
Because the graph can get very cluttered when many model semivariograms are
displayed at once, the Latin-Hypercube:Individual Display option is available (pop-up
dialog in Figure 9.15). It allows the user to examine each model semivariogram
individually (Figure 9.16). This is also convenient for examining models which are
marked invalid. If the Solution # for the model is known (unlikely), it can be entered
directly, or the user can step through the models individually by using the Next and
Previous buttons. For each model, the user can mark it as Valid, Invalid (Discard), or
yet to be determined (?).
 Figure 9.15
Figure 9.15
 Figure 9.16
Figure 9.16
- NOTE: Only Valid models are saved.
When the dialog is destroyed, the full series of model semivariograms is redisplayed.
Latin-Hypercube:Save saves the numerical model results of the Latin-Hypercube
to a data file readable by sisim (Chapter 14 ) (See the Output Data File Format section for
file format details). If the output file has already been named, the old file is overwritten
and the new data is saved. If no file has been named, the user is requested for a
filename (*.lhc extension) with a pop-up dialog similar to that shown in Figure 5.2.
Latin-Hypercube:Save as creates a pop-up dialog similar to that shown in Figure 5.2 requesting an output file name for the Latin-Hypercube sampled model
semivariograms (default extension = *.lhc). Once the file name is selected, the data is
saved.
The Latin-Hypercube:Calculate menu-bar option calculates the latin-Hypercube
sampled model semivariograms based on the parameters define in the Latin-
Hypercube:Modify dialog.
[TOP] [SYNTAX]
Series allows the user to load in a series of experimental semivariograms. These
will usually be calculated from a series of indicator simulations (See Chapter 14 , sisim).
This is a useful tool for viewing the ergodic fluctuations of the simulation results ()
versus the original input/model semivariogram used to calculate them. In theory there
should be variation in the simulation grid experimental semivariograms, but they
should, in sum, reflect the initial model semivariogram they were all calculated from.
This tool can be used to evaluate how well this assumption is honored.
Series:Display creates the pop-up dialog shown in Figure 9.17. This dialog is
used to control what information about the experimental semivariograms is displayed
on the graph. Under the default configuration, each semivariogram is drawn, along
with the mean g(h) value for every lag, and the maximum extents of the sampled g(h)
values (Figure 9.18a). Various details can be turned on or off. In Figure 9.18b, only the
mean lag g(h) values and the g(h) extents are display. In Figure 18c, the 95% confidence
intervals of the lag g(h) values are added. In Figure 18d, only the g(h) extents, and an
individual model are displayed.
 Figure 9.17
Figure 9.17
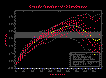 Figure 9.18a;
Figure 9.18a;
 Figure 9.18b;
Figure 9.18b;
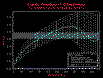 Figure 9.18c; and
Figure 9.18c; and
 Figure 9.18d.
Figure 9.18d.
Series:Parameters creates the pop-up dialog shown in Figure 9.19. This dialog is
used to control what experimental semivariograms are loaded into variofit. The files
loaded need to use the following naming convention:
- prefix.#.suffix
 Figure 9.19
Figure 9.19
An example series might be:
- sample.1.gam
sample.2.gam
sample.3.gam
sample.4.gam
If there are models missing between the first and last selected, a warning message will
be printed in the log/status window. The fact that there is a file missing will not effect
the calculated statistics. The statistics are based only on the models successfully
loaded.
[TOP] [SYNTAX]
Graph allows the user to specify various attributes about the appearance of the
graph. Attributes about the graph Border, Error-bar Styles, Fonts, Labels, Legends,
Mesh, and Line Styles.
Graph:Border is described in Chapter 5 in the Graph:Border section (Figure 5.9).
Error-Bar Style controls the display characteristics of the error bars. Error bars
only exist when a jackknifed calculation has been done. The Error-Bar Style dialog is
shown in Figure 7.22. The Error-Bar Style settings are very similar to the settings
available in the Style: Set Line Attributes dialog as described in Chapter 5 (Figure 5.14).
The only difference in the dialogs is the option to set the extent of the cross-width on
the error-bars.
Graph:Fonts is described in Chapter 5 in the Graph:Fonts section (Figures 5.10 and 5.11).
Graph:Labels is described in Chapter 5 in the Graph:Labels section (Figure 5.12).
Data Parameters:
Graph:Legends:Data Parameters is described in Chapter 8 in the
Graph:Legends:Data Parameters section (Figure 7.23).
Model Parameters:
Graph:Legends:Model Parameters dialog is shown in Figure 9.20. It is very
similar to the Graph:Legends:Data Parameters dialog (Figure 7.23), except this option
controls if and where the model semivariogram parameters will be displayed on the
graph. An example is shown in Figure 9.1.
 Figure 9.20
Figure 9.20
Graph:Mesh is described in Chapter 5 in the Graph:Mesh section (Figure 5.13).
Graph:Line Styles is described in Chapter 5 in the Graph:Style section (Figure 5.14).
[TOP] [SYNTAX]
The Log menu option is supplied to allow the user to save, view, or print all text
which has been written to the log/status window by the program or added by the user (The log window is also a simple text editor). The options include View Log, Save, Save
as, Clear, and Print. View Log, Clear, Save, and Save as are similar in operation to the
menu options under File described above.
[TOP] [SYNTAX]
Plot:Now and Plot:Refresh are described in Chapter 5 in the Plot:Now and
Plot:Refresh sections.
[TOP] [SYNTAX]
Help works exactly as explained in Chapter 5 (plotgraph, Figure 5.15) Help
section.
[TOP] [SYNTAX]
Using variofit is not as straight forward as many of the other UNCERT modules.
This package assumes that the user is familiar with the concepts of experimental and
model semivariograms, and the basics of fitting model semivariograms. If the user is
familiar with these concepts, variofit is relatively easy to use correctly. Before taking
advantage of the jackknifing and latin-hypercube options, though the user should be
thoroughly familiar with the appropriate Mathematical Sections for vario (Chapter 8),
variofit, issim (Chapter 14 ).
The difficulty in using this package comes from generating the model
semivariogram correctly. Once this has been done, the appearance of the graph can be
modified using many of the same tools described for plotgraph in Chapter 5 .
There are three methods to load a file in variofit. The first is to execute variofit
from the UNIX prompt and open the file from the menu, the second is to pass the file as
a command line argument, and the third is to define the file name in the program
preference file. To open a file from the main menu, execute variofit from the UNIX
prompt:
- > variofit
Once in the application, select the File:Open menu option. The pop-up dialog shown in
Figure 5.2 will appear. Select the desired file. Once a file has been selected the graph
of the data will be drawn. To open a file from the command line, enter at the UNIX
prompt:
- > variofit [optional arguments] filename
For Example:
- > variofit -ymin 0.0 clark.gam
- NOTE: This show Figure 9.3 although other variables then those passed on the
command line here were defined. These variables could have been set using the
menus, or using a preference file (Appendix C). A preference file is used to
define user preferred variable default values. Every time variofit runs, it
searches the current working directory for the file variofit.prf. If it exists, variofit
reads the file and sets the variables as specified. This is the third way to open a
file, because one of the arguments in the preference file is the name of the graph
file.
Once an experimental semivariogram is loaded (non-jackknifed), it is a simple
matter to automatically fit a rough model semivariogram to the data. To display the
available options, select the Regression:Parameters menu-bar option.
To fit a model semivariogram to the experimental semivariogram in Figure 9.3
(clark.dat), an initial guess might be to fit a single-nested spherical model (Select a
Spherical model for nest 1) to the data set. The sill appears to approximately equal the
data set variance; for this reasons, using the Variance LS Solution Method may be
appropriate. When Regression:Calculate is pressed the model semivariogram in Figure 9.21a is generated.
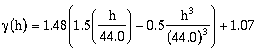
This model is a reasonable approximation based on the constraints used, but it is not a
good fit. The calculated nugget of 1.07 is much larger then one would expect.
Extending the slope of the first few lags suggests a value closer to 0.4! The model, also,
in zones tends to consistently over- or under-estimate the experimental semivariogram
lag values. A better model selection may be a two-nested spherical structure (Select a
Spherical model for nest 2). For nests one and two, select the Spherical model type.
When the model is recalculated (Press Regression:Parameters), the model shown in
Figure 9.21b is a better fit.
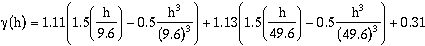
This, model honors the early portion of the experimental semivariogram more
accurately then the single nested model and is therefore a better fit. It does still tend to
overestimate the variance over the 15 to 45 meter lag distance. Using the automatic
fitting algorithm, however, this is about the best that will be done.
 Figure 9.21a, and
Figure 9.21a, and
 Figure 9.21b
Figure 9.21b
Note that this automatic model fitting process is not completely automatic. The
modeler is responsible for selecting the appropriate model structure, the algorithm only
fits a curve based upon set constraints. If the constraints are poorly defined, the model
fit will also be poor.
To improve upon the automatic model fit discussed above (Figure 9.21b), the
user will have to manually adjust the model parameters. First set the Calculation
Method to Manual. By examining the model and experimental semivariograms in Figure 9.21b it appears that the transition from the first to the second nest may be starting at
slightly to short of a lag (Break is above all surrounding lag points) and the range for
nest two maybe short also (Most lags point between 15 and 45 meters fall below model
curve). By increasing the range values to 11m and 55m, and slightly adjusting the
variance components for each lag, a good fit is modeled (Figure 9.21c).
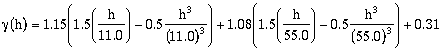
Although this model could be further refined, there is probably little reason to do so;
with the kriging process, there would probably be little difference between a map
generated with this model and a more refined model. Practically , there would probably
be little difference between the model shown in Figure 9.21b and Figure 9.21c.
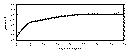 Figure 9.21c
Figure 9.21c
Fitting a series of Latin-Hypercube sampled model semivariograms to a series of
jackknifed experimental semivariograms is a simpler process then fitting a single model
semivariogram, because the lag values do not have to be honored, only the range of
uncertainty has to be honored. This process is also highly automated (There is no
manual mode), and the modeler is free only to set some constraints, and accept or
reject calculated model semivariograms.
As an example, load (File:Open) the experimental jackknifed semivariogram file
well.jack.gam. This series of experimental semivariograms describes the uncertainty in
the spatial distribution of materials based on the wells shown in Figures 1.3 and 1.4.
The data set (well.jack.gam) was calculated by vario ( Chapter 8 example). Once the file
is loaded, the jackknifed experimental semivariograms and their uncertainty will be
displayed (Figure 9.22a).
 Figure 9.22a
Figure 9.22a
The Latin-Hypercube technique allows the modeler to select a limited number of
model semivariograms which cover the range of solutions which honor the confidence
intervals of jackknifed experimental semivariograms and the modelers opinion of
reasonable values. After loading the file, the modeler must specify how to sample the
jackknifed experimental semivariograms to generate model semivariograms. For this
data set there, is definitely not enough data to suggest a two-nested structure. Model
this data set with only one nest. Select a Spherical model for nest one. A spherical
model is selected because it has given reasonable results for similar conditions. Based
on the data available, it would be difficult to argue one model was better than another.
Because the selection criteria for range needs work, reset the Minimum Range to 10.0
(data implies pure-nugget effect possible, a 0.0 range though is felt highly unlikely
(expert opinion)), and the Maximum Range to 230.0 (Confidence interval of third lag
suggests range could be greater then 230 feet but it is unlikely to be much longer).
Setting these limits sets reasonable ranges for the Range variable. Once these parameters are set, select Latin-Hypercube:Calculate. Sixteen model semivariograms
will be generated and displayed (4 Range Divisions * 1 Variance Division * 4 Nugget
Divisions * 1 Solution per Division). Every time Latin-Hypercube:Calculate is pressed,
using these parameters these exact 16 model semivariograms will be calculated. To
generate a different set of random sampled model semivariograms, change the
Randomize Seed value to another integer value (Set to 10, it was 1 (default value)). If
this is done, a new series of model semivariograms will be calculated (Figure 9.22b).
Note that in this series, two models are drawn with dashed red lines. This indicates
that these models do not honor the uncertainty limits of the experimental
semivariograms and should not be used for future calculations. To save these valid
models, Select the Latin-Hypercube:Save menu option and supply an appropriate file
name. This file can be used for input for the sisim module (Chapter 14 ).
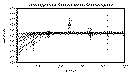 Figure 9.22b
Figure 9.22b
- NOTE: For different levels of uncertainty, the confidence intervals can be reduced
or expanded in the Jackknife:Modify dialog.
[TOP]
In many cases it is more convenient to run the application completely from the
command line, or at least pass some parameter values in from the command line. The
options listed below allow the user to accomplish almost anything that is possible from
within the X-windows application from the command line (adding lines from different
files is not currently supported). This feature can be useful when the user does not
have a X-windows/Motif terminal available, or when many graphs need to be processed
quickly, and the operation can be completed in batch mode without user interaction.
Syntax:
- Meaning of flag symbols:
- # = integer
#.# = float
" " = character string.
{} = variable is an array. Values must be seperated by a ',' and no spaces are allowed. Do not use the "{ }" symbols on the command line.
NOTES:- 1) All parameters in [] brackets are optional.
2) Quotes must be used around character strings.
3) Filename, if given, must be listed last.
4) If no default is given, the feature is not currently supported on command line.
If no entry is required for flag, flag command executed.
Flag Definitions:
-
| -am | = |
active experimental model (for calculation) | default = 1 |
| -cs | = |
calculation steps for automatic solution | default = 100 |
| -er | = |
ending range | default = 0.0 |
| -esp | = |
exageration scale priority | default = 0 |
| |
0
1 |
=
= |
favor y-exageration scale (-ys)
favor x/y ratio |
|
| -fnt1 | = |
main title font | default = Helvetica-Bold |
| -fnt2 | = |
secondary title font | default = Helvetica-Bold |
| -fnt3 | = |
axes label font | default = Helvetica |
| -fnt4 | = |
division font | default = Helvetica |
| -fnt5 | = |
annotation font | default = Helvetica |
| -fnt6 | = |
mouse position font | default = Helvetica |
| -fnts1 | = |
main title font size | default = 24.0 |
| -fnts2 | = |
main title font size | default = 15.0 |
| -fnts3 | = |
main title font size | default = 15.0 |
| -fnts4 | = |
main title font size | default = 12.0 |
| -fnts5 | = |
main title font size | default = 10.0 |
| -fnts6 | = |
main title font size | default = 12.0 |
| -help | = |
give this help menu | |
| -jcl | = |
jackknife error-bar/band confidence level | default = 90.0% |
| -jebd | = |
plot jackknife error-bands (lag variance) | default = 1 |
| |
|
| -jebr | = |
plot jackknife error-barss (g(h) variance) | default = 1 |
| |
|
| -jev | = |
plot jackknife error variance bars | default = 1 |
| |
|
| -jir | = |
plot intermediate jackknife results | default = 1 |
| |
|
| -jjr | = |
plot full unjackknifed semivariogram | default = 1 |
| |
|
| -lc {} | = |
line color | default = variable |
| |
0
1
2
3
4
5
6
7 |
=
=
=
=
=
=
=
= |
Black
White
Red
Green
Blue
Magenta
Yellow
Cyan |
|
| -lgf | = |
log file name | defalut = "log.dat" |
| -lgmp | = |
legend parameter position | default = 1 |
| |
0
1
2
3 |
=
=
=
= |
Top left
Top right
Bottom left
Bottom right |
|
| -lgms | = |
legend model displayed | default = 1 |
| |
|
| -lgpp | = |
legend parameter position | default = 1 |
| |
0
1
2
3 |
=
=
=
= |
Top left
Top right
Bottom left
Bottom right |
|
| -lhck | = |
check latin-hypercube samples in range | default = 1 |
| |
|
| -lhmax1 | = |
latin-hypercube maximum range | default = max. lag |
| -lhmax2 | = |
latin-hypercube maximum range | default = max. variance |
| -lhmax3 | = |
latin-hypercube maximum range | default = max. variance |
| -lhmax1 | = |
latin-hypercube maximum range | default = min. lag |
| -lhmin2 | = |
latin-hypercube maximum range | default = min. variance |
| -lhmin3 | = |
latin-hypercube maximum range | default = min. variance |
| -lhmt1 | = |
latin-hypercube nest #1 model structure | default = 3 |
| |
0
1
2
3
4
5
6
7 |
=
=
=
=
=
=
=
= |
No Model (Not available for nest #1)
Linear
Linear With Sill
Spherical
Exponential
Logarithmic
Power (NOT INSTALLED)
Gaussian |
|
| -lhmt2 | = |
latin-hypercube nest #2 model structure (See -lhmt1) | default = 3 |
| -lhout | = |
latin-hypercube output file | default = "junk.lhc" |
| -lhrj | = |
draw only latin-hypercube samples in range | default = 0 |
| |
|
| -lhseed | = |
latin-hypercube random seed | default = 1.0 |
| -lhseg1 | = |
latin-hypercube range divisions | default = 4 |
| -lhseg2 | = |
latin-hypercube C divisions | default = 1 |
| -lhseg3 | = |
latin-hypercube nugget divisions | default = 4 |
| -lhsel | = |
latin-hypercube range selection | default = 1 |
| |
0
1 |
=
= |
equally spaced
ramdom |
|
| -lhspd2 | = |
latin-hypercube solutions per division | default = 1 |
| -lpbm | = |
page bottom margin | default = 1.5 |
| -lpc | = |
number of copies to print | default = 1 |
| -lpd | = |
print destination | default = 0 |
| |
|
| -lpf | = |
print filename | default = "junk.ps" |
| -lph | = |
print header page | default = 0 |
| |
|
| -lplm | = |
page left margin | default = 1.5 |
| -lpo | = |
print orientation | default = 0 |
| |
0
1 |
=
= |
Portrait
Landscape |
|
| -lppsext | = |
search extention for postscript files | default = "*.ps" |
| -lpq | = |
print queue | default = "ps" |
| -lpr | = |
print file at specified orientations | |
| -lprm | = |
page right margin | default = 1.0 |
| -lps | = |
print output | default = 0 |
| |
0
1 |
=
= |
Black & white
Color |
|
| -lptm | = |
page top margin | default = 1.5 |
| -lsfl {} | = |
fill line symbol | default = 0 |
| |
|
| -lsc {} | = |
line symbol color | default = variable |
| |
1
2
3
4
5
6
7 |
=
=
=
=
=
=
= |
Black
White
Red
Green
Blue
Magenta
Yellow
Cyan |
|
| -lssz {} | = |
line synbol size | default = 9.0 |
| -lsty {} | = |
line symbol type | default = 0 |
| |
-1
0
1
2
3
4 |
=
=
=
=
=
= |
No Symbol
Circle
Cross
Diamond
Square
X |
|
| -ltk {} | = |
line thickness | default = 1.0 |
| -lty {} | = |
line type | default = 0 |
| |
-1
0
1
2 |
=
=
=
= |
No Line
Solid
Dashed
Double Dashed |
|
| -m1 | = |
model types nest #1 | default = 3 |
| |
0
1
2
3
4
5
6
7 |
=
=
=
=
=
=
=
= |
No Model (Not available for nest #1)
Linear
Linear With Sill
Spherical
Exponential
Logarithmic
Power (NOT INSTALLED)
Gaussian |
|
| -m2 | = |
model types nest #2 (See -m1) | default = 0 |
| -m3 | = |
model types nest #3 (See -m1) | default = 0 |
| -m4 | = |
model types nest #4 (See -m1) | default = 0 |
| -md | = |
dash mesh | default = 0 |
| |
|
| -mox | = |
X mesh origin | default = 0.0 |
| -moy | = |
Y mesh origin | default = 0.0 |
| -ms | = |
use mesh | default = 0 |
| |
|
| -mx | = |
X mesh frequency | default = 1/10 DX |
| -my | = |
Y mesh frequency | default = 1/10 DY |
| -ng | = |
nugget | default = 0.0 |
| -out | = |
output *.out filename | defalut = "junk.out" |
| -prf | = |
preference file name | defalut = "variofit.prf" |
| -r1 | = |
range for nest #1 | default = 0.0 |
| -r2 | = |
range for nest #1 | default = 0.0 |
| -r3 | = |
range for nest #1 | default = 0.0 |
| -r4 | = |
range for nest #1 | default = 0.0 |
| -rfh | = |
screen refresh | default = 0 |
| |
0
1 |
=
= |
On exposure
On update |
|
| -run | = |
run and calcualte variofit without X-interface | |
| -s1 | = |
sill for nest #1 | default = 0.0 |
| -s2 | = |
sill for nest #1 | default = 0.0 |
| -s3 | = |
sill for nest #1 | default = 0.0 |
| -s4 | = |
sill for nest #1 | default = 0.0 |
| -scl | = |
series confidence level | default = 95.0 |
| -sdc | = |
draw series lag confidence bars | default = 1 |
| |
|
| -sdm | = |
draw series lag mean | default = 1 |
| |
|
| -sdx | = |
draw series lag extents | default = 1 |
| |
|
| -sex | = |
series extention | default = ".gam" |
| -sp | = |
series prefix | default = " " |
| -sr | = |
search range | default = Max. diagonal |
| -sse | = |
ending series | default = 1 |
| -sss | = |
starting series | default = 1 |
| -st | = |
automatic solution type fitting algorithm | default = 4 |
| |
0
1
2
3
4 |
=
=
=
=
= |
Least-Squares
Nugget Least-Squares
Variance Least-Squares
Nugget & Variance Least-Squares
Weighted Least-Squares |
|
| -sttl | = |
Secondary title | default = " " |
| -sv | = |
solver method | default = 0 |
| |
|
| -svm | = |
experimental semivariogram read from file | default = 1 |
| |
|
| -ttl | = |
Main title | default = Filename |
| -xfmt | = |
Number of decimal places for X-axis | default = ".2f" |
| -xlabel | = |
X-axis label | default = "X" |
| -xmax | = |
Graph X-maximum | default = Data Maximum |
| -xmin | = |
Graph X-minimum | default = Data Minimum |
| -xMt | = |
X main tic frequency | default = 1/10 DX |
| -xmt | = |
Number of minor X tics | default = 5 |
| -xto | = |
X axis label origin | default = 0.0 |
| -xy | = |
xy ratio | default = 1.5 |
| -yfmt | = |
Number of decimal places for X-axis | default = ".2f" |
| -ylabel | = |
X-axis label | default = "Y" |
| -ymax | = |
Graph Y-maximum | default = Data Maximum |
| -ymin | = |
Graph Y-minimum | default = Data Minimum |
| -yMt | = |
X main tic frequency | default = 1/10 DY |
| -ymt | = |
Number of minor Y tics | default = 5 |
| -ys | = |
Y-axis exageration relative to X-axis | default = Calculated |
| -yto | = |
X axis label origin | default = 0.0 |
An example command might be (typed on one line):
- variofit -xmin 0.0 -ymin 0.0 -ymax 12.0 -md 1 -ms 1 -mx 1.0 -my 1.0 -xMt 1.0 -yMt
1.0 -ttl "Semivariogram of Elevation Data" -sttl "UNCERT variofit module" -xlabel
"distance (feet)" -ylabel "gamma h" -xfmt ".1f" -yfmt ".1f" -esp 1 -xy 1.0 water.out
[TOP]
Variofit can read two data file formats; one is specified for single experimental
semivariograms, and the other is for multiple jackknifed experimental semivariograms.
The user does not have to explicitly tell the program what format the data file is; the
program can determine this based of the file format.
The single model format name is misleading. Multiple models may be stored
with this format, but each model is saved with its own set of model parameters used to
create it (Various search parameters including the lag, search direction, bandwitdhs,
etc.) The format is outlined below.
- SEMIVARIOGRAM #[Model Number][/][Number of Models in File]
- for (i=1;i<=# of Models)
- [Lags in Model]
for (j=1;j<=# of Lags)
- [Minimum Lag][Maximum Lag][Average Lag][
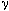 (h)][Pairs]
(h)][Pairs]
end for
[Data Set Variance]
[Data Set Mean]
[Calculation Type: 0 = Point, 1 = Gridded]
if (calculation type == Point)
- [Lag]
[Search Direction]
[Plunge]
[Horizontal Bandwidth]
[Vertical bandwidth]
else
- [X Step increment]
[Y Step increment]
[Z Step increment]
endif
[Spatial Equation: 0 = Semivariogram, 1 = Cross-Semivariogram, ... (See
-set flag in vario (Chapter 8))]
if (spatial equation == Indicator or Soft-Indicator Covariance)
- [Low Cutoff]
[High Cutoff]
endif
[Data Set File Name]
if (calculation type == Point)
- [X Column]
[Y Column]
[Z Column]
[Head Column]
if (spatial equation == Covariance, Cross-Semivariogram,
Correlogram, or Soft Indicator Covariance)
- [Tail Column]
endif
endif
end for
If there is only one model, the data from Data Set Varaince down can be ignored. For an
example file, see data file water.3.gam.
The jackknife model format stores all the jackknifed experimental
semivariograms and details about the variance over each lag interval. The format is
outlined below:
- JACKKNIFED SEMIVARIOGRAM
[Data Set Mean] [Data Set Variance]
[# of Lags]
for (i=1;i<=# of Lags)
- [Lag #] [Mean of Lag] [Lag Standard Deviation] [�.7;(h) Standard Deviation]
end for
[# of Experimental Semivariograms]
for (i=1;i<=# of Semivariogram)
- [# of Lags for this Semivariogram]
for (j=1;j<=# of Lags)
- [Minimum Lag][Maximum Lag][Average Lag][
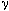 (h)][Pairs]
(h)][Pairs]
end for
end for
For an example file, see data file well.jack.gam.
[TOP]
For variofit data files, there are two output file formats, one for single model
semivariograms and one for multiple Latin-Hypercube sampled model semivariograms.
The output for a single model semivariogram is a commented header describing
the parameters for each nest of the model semivariogram followed by three line
descriptions (using MULTIPLE LINE format, plotgraph, Chapter 8) specifying the data
set variance, the experimental semivariogram and the model semivariogram. The
header for the model semivariogram shown in Figure 9.3 is:
!-------------------------------------
!Model Segment 1: Spherical
! Range = 9.600000
! Sill = 1.109714
!
!Model Segment 2: Spherical
! Range = 49.600000
! Sill = 1.133128
!
!Nugget = 0.307158
!
!Variance = 2.550000
!Mean = 3.041230
!
!Sum squared error = 0.169223
!-------------------------------------
MULTIPLE LINES
3
2 1 5 1.0 -1 5 5.0 1 Variance
0.00000 2.55000
80.00000 2.55000
0 -1 2 1.0 0 2 5.0 1 Variance
200 0 4 3.0 -1 4 5.0 1 Variance
0.00000 0.30716
0.40000 0.39018
.
.
.
basically this file can be used as a reference to lookup model parameters, and it a valid
graph file for plotgraph (Chapter 8).
The output file for Latin-Hypercube sampled models is not a valid graph file like
the output file for a single model semivariogram. This file is used for input to sisim
(Chapter 14 ) and describes each model semivariogram. The format is (Values in []
required entries; indentation shown not required - used to indicate loops):
- [# of Thresholds]
for (i=1;i<=# thresholds)
- [Threshold] [Prior CDF]
[# of models for threshold]
for (j=1;j<=# of models)
- [# of nests] [nugget]
for (k=1;k<=# of nests)
- [Model Type] [Range] [Sill] [X-Anis] [Y-Anis]
end for
[Major X-Y Search Direction] [Plunge Angle]
end for
end for
For an example file, see data file well.aniso.2.lhc.
[TOP]
See Chapter 8 (vario) Mathematics Section on semivariogram calculations.
Because of the difficulty inexperienced geostatisticians have with fitting a model
to an experimental semivariogram, and the difficulty experienced geostatisticians have
with fitting nested models to an experimental semivariogram, an automated model
fitting procedure is included in the software. The procedure is based on a least-squares
minimization approach and is used to minimize the sum of squared error terms:
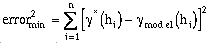 (9-1)
(9-1)
This process is not perfect, but it does yield reasonable results and it can give the
experienced user a good starting equation when fitting complicated nested models.
In the case of a Spherical Model with a nugget and one sill, the model
semivariogram is defined as (Journel and Huijbregts, 1978):
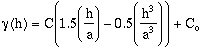 ; for h < h <= a. (9-2)
; for h < h <= a. (9-2)
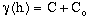 ; for h > a. (9-3)
; for h > a. (9-3)
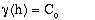 ; for h = 0. (9-4)
; for h = 0. (9-4)
- C = point variance.
Co = nugget or random variance.
a = range.
h = sample distance.
The estimation error is defined as:
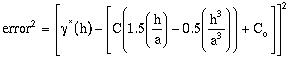 ; for h <= a (9-5)
; for h <= a (9-5)
where,
-
 = experimental semivariogram value at h,
= experimental semivariogram value at h,
and defining:
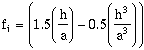 (9-6)
(9-6)
then the squared error at h is:
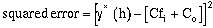 (9-7)
(9-7)
Squaring the error and expanding equation (9-7) over the number of samples, and
summing from i = 1 to n (n = number of sample pairs):
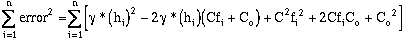 (9-8)
(9-8)
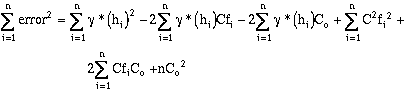 (9-9)
(9-9)
Equation (9-9) is minimized by setting its first two derivatives with respect to C and Co
equal to zero:
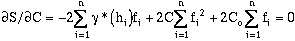
therefore
 (9-10)
(9-10)
and
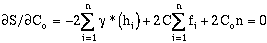
therefore
 (9-11)
(9-11)
Using a matrix form, the two unknowns, C and Co, can be calculated by:
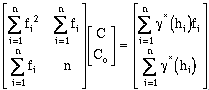 (9-12)
(9-12)
This equation is in the form:
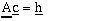
Where

is the equation to be solved. Once the combination of C and Co which minimize the
solution are determined, the sum of squared errors is calculated, and a new range (a) is
selected. This is done iteratively until the range (a) which yields the minimum error is
found. Generally, it is sufficient to evaluate "a" to plus or minus 1/100th of the
maximum distance between data points (Instead of using the maximum distance across
the data set, another distance may be more appropriate and yield more stable
estimates). The form of the solution matrices in equation (9-12) are the same regardless
of the type model that is fit to the data. The only difference is that the term fi, is
evaluated using the appropriate model. These are defined as follows (Journel and
Huijbregts, 1978):
- Exponential Model:
 (9-13a)
(9-13a)
Gaussian Model:
 (9-13b)
(9-13b)
Logarithmic Model:
 (9-13c)
(9-13c)
Linear Model:
 (9-13d)
(9-13d)
Linear With Sill Model:
 (9-13e)
(9-13e)
Spherical Model:
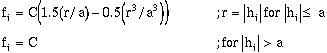 (9-13f)
(9-13f)
where for the linear models, m is a slope term. NOTE: In the future Power and Hole
Effect models will be included in the software.
- Power Model:
 (9-13g)
(9-13g)
When describing the experimental semivariogram, often a nugget and a single point
variance (i.e. one sill) term is inadequate. Models that fit such data have what is called
a nested structure, and may be described by several point variance terms (i.e., C0, C1,
C2,...). So, for a two-nested semivariogram:
 (9-14)
(9-14)
where:
- C0 = nugget.
C1 = point variance of the first nest with range a1.
C2 = point variance of the second nest with range a2, where a2 >= a1.
di, eii = an experimental model, as f used in equations 9-13a-g. Note that
these models may be of different types.
Under this arrangement, three terms, C0, C1, and C2, must be determined. The
solution matrix is:
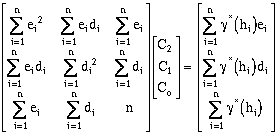 (9-15)
(9-15)
For a model with three nests, four terms, C0, C1, C2, and C3 must be determined. The
format is similar:
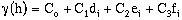 (9-16)
(9-16)
where
- C0 = nugget.
C1 = point variance of the first nest with range a1.
C2 = point variance of the second nest with range a2, where a2 >= a1.
C3 = point variance of the second nest with range a3, where a3 >= a2 >= a1.
di, ei, fi = n experimental model, as f used in equations 9.13a-g. Note that
these models may be of different types.
and the solution matrix is:
 (9-17)
(9-17)
To demonstrate how these matrices were developed, the final case developed where four
nests are used will be derived. In this case there are nine unknowns, the five point
variance terms C0, C1, C2, C3, and C4, and four ranges. In this case, as in the previous
cases the point variance terms are determined explicitly, but the four range terms are
determined through an iterative process. For the system where:
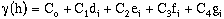 (9-18)
(9-18)
where
- C0 = nugget.
C1 = point variance of the first nest with range a1.
C2 = point variance of the second nest with range a2, where a2 >= a1.
C3 = point variance of the third nest with range a3, where a3 >= a2 >= a1.
C4 = point variance of the fourth nest with range a4, where a4 >= a3 >= a2 >= a1.
di, ei, fi, gi = an experimental model, as f is used in equations 9-13a-g. These
models may be of different types.
The error at a specific lag distance is:
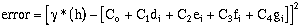 (9-19)
(9-19)
Expanding the error terms and minimizing yields (equations have been divided by 2.0):
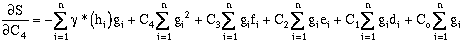


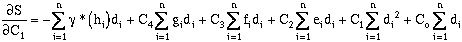
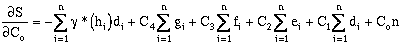
where:

rearranging:
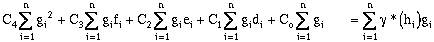 (9-20)
(9-20)
 (9-21)
(9-21)
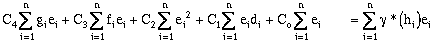 (9-22)
(9-22)
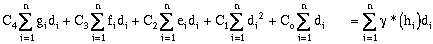 (9-23)
(9-23)
 (9-24)
(9-24)
Yielding the solution matrix:
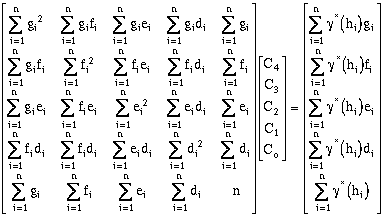 (9-25)
(9-25)
The approach describe above, explains the process for calculating the least-squares
"best-fit" curve for a given set of range values, a1, a2, a3, and a4, where the nugget (Co)
and variance terms (C1, C2, C3, and C4) are unknown.
Modified Least-Squares Regressions:
In some cases, the user may by able to specify further information about the
experimental semivariogram, or about expected results for the model semivariogram. A
common modeling method is to assume the model sill equals the data set variance, i.e.:
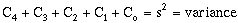 (9-26)
(9-26)
Another possible constraint is to assume that the nugget (Co) equals the y-intercept of a
line drawn through the shortest two lag points in the experimental semivariogram.
Here the nugget equals:
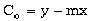 (9-27)
(9-27)
where x and y are the range and 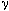 (h) coordinates of the first lag point, and m is the
slope of the line between the shortest two lag points. This isn't necessarily a good
assumption, but on a practical basis, it can improve the model solution in some cases.
Another alternative is to weight each lag point by the number of data pairs that went
into defining that point, the theory being that a point represented by many data points
should be honored more closely then a lag point only based on a few sample pairs.
Once the statistical distribution of experimental semivariograms has been
calculated, semivariograms can be fit through the zone defined by the error-bars. The
objective is not to make a single best estimate of the character of the subsurface (i.e. a
single semivariogram), rather the objective is to select model semivariograms
representative of the range of possible conditions at the site. This range of
semivariograms is used with the original data to conduct indicator kriging and
stochastic simulation to generate multiple interpretations of the subsurface. One
approach is to use Monte-Carlo techniques and randomly select 100 model
semivariograms that fall within the range of reasonable solutions. This might appear
reasonable, but expert opinion of conditions at the site may indicate that models
generated with nuggets approximately equal to the sill or ranges near zero are
unreasonable or unrealistic, even though the jackknifed experimental semivariogram
indicates such semivariogram models of the site are possible interpretations.
(h) coordinates of the first lag point, and m is the
slope of the line between the shortest two lag points. This isn't necessarily a good
assumption, but on a practical basis, it can improve the model solution in some cases.
Another alternative is to weight each lag point by the number of data pairs that went
into defining that point, the theory being that a point represented by many data points
should be honored more closely then a lag point only based on a few sample pairs.
Once the statistical distribution of experimental semivariograms has been
calculated, semivariograms can be fit through the zone defined by the error-bars. The
objective is not to make a single best estimate of the character of the subsurface (i.e. a
single semivariogram), rather the objective is to select model semivariograms
representative of the range of possible conditions at the site. This range of
semivariograms is used with the original data to conduct indicator kriging and
stochastic simulation to generate multiple interpretations of the subsurface. One
approach is to use Monte-Carlo techniques and randomly select 100 model
semivariograms that fall within the range of reasonable solutions. This might appear
reasonable, but expert opinion of conditions at the site may indicate that models
generated with nuggets approximately equal to the sill or ranges near zero are
unreasonable or unrealistic, even though the jackknifed experimental semivariogram
indicates such semivariogram models of the site are possible interpretations.
An alternative approach to random selection of a large number of possible
semivariogram models is to use latin-hypercube sampling. This reduces the number of
simulations required to insure that the "flavor" of all alternatives is addressed (McKay et
al, 1979).
This approach can yield a daunting number of simulations, many of which will
bear little resemblance to one another if the data set is small. Such a situation results
in the obvious conclusion that some data sets provide so little information about a site
that more data should be collected before further assessment is undertaken. If the data
are more abundant, the range of possible models will be constrained, and the simulated
models may represent a modest range of possible subsurface interpretations. If the
jackknifed semivariogram has small error-bars, the entire process of using a variety of
semivariograms for simulation of one site can be omitted because the process is not
likely to indicate a larger uncertainty associated with the interpretation of such well
characterized sites.
Recall that the objective of this approach is not to make a single best estimate of
the subsurface interpretation, but to evaluate the possible range of subsurface
character based on available data. From a purely mathematical approach this may be
computationally intractable, however incorporation of expert opinion into the process
makes it possible to limit the reasonable alternatives.
[TOP]
Burden, R.L. and J.D. Faires, 1985, Numerical Analysis, Third Edition, Prindle, Weber, and Schmidt, Boston, pp 342-353.
Clark, I., 1979, Practical Geostatistics, Elsevier Applied Science, London and New York.
Journel, A.G. and Ch. J. Huijbregts, 1978, Mining Geostatistics, Academic Press, London.
McKay M.D., R.J. Beckman and W.J. Conover. 1979. "A Comparison of Three Methods
for Searching of Input Variables in the Analysis of Output From a Computer Code",
Technometrics. Vol. 21, No. 2, pp 239-245.
Wingle, W.L., and E.P. Poeter, 1993, Evaluating Uncertainty Associated with
Semivariograms Applied to Site Characterization. Ground Water, Vol. 31, No. 5, pp
725-734.
[TOP]
Table of Contents
Previous Chapter
Beginning of this Chapter
Next Chapter


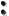

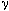 (h) lag values. Once the nugget is determined
the standard least-squares approach used above is applied.
(h) lag values. Once the nugget is determined
the standard least-squares approach used above is applied.
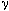 (h) value (i.e., this method presumes that it is more important
to honor a data point representing 500 data pairs than one representing 3 data
pairs). The previous four methods do not consider the number of data pairs,
therefore they weight all lag values equally.
(h) value (i.e., this method presumes that it is more important
to honor a data point representing 500 data pairs than one representing 3 data
pairs). The previous four methods do not consider the number of data pairs,
therefore they weight all lag values equally.
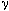 (h) value for
each lag are drawn. Draw Error-Bands determines whether the horizontal error-bars
representing the confidence limits of the lag distance are drawn. Draw Range of
Jackknifed Variance's: because each jackknifed experimental semivariogram is based
on a slightly different set of data points the variance for each solution is also different;
this option determines whether this variation is drawn. Finally, the confidence level for
the error-bars can be specified; 95% is the default.
(h) value for
each lag are drawn. Draw Error-Bands determines whether the horizontal error-bars
representing the confidence limits of the lag distance are drawn. Draw Range of
Jackknifed Variance's: because each jackknifed experimental semivariogram is based
on a slightly different set of data points the variance for each solution is also different;
this option determines whether this variation is drawn. Finally, the confidence level for
the error-bars can be specified; 95% is the default.
Figure 9.13a,
Figure 9.13b, and
Figure 9.13c
Figure 9.18a;
Figure 9.18b;
Figure 9.18c; and
Figure 9.18d.
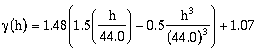
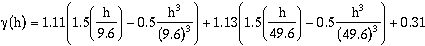
Figure 9.21a, and
Figure 9.21b
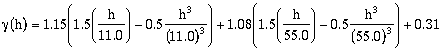
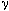 (h)][Pairs]
(h)][Pairs]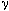 (h)][Pairs]
(h)][Pairs]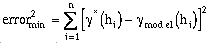 (9-1)
(9-1)
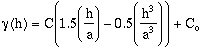 ; for h < h <= a. (9-2)
; for h < h <= a. (9-2)
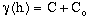 ; for h > a. (9-3)
; for h > a. (9-3)
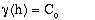 ; for h = 0. (9-4)
; for h = 0. (9-4)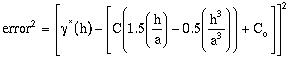 ; for h <= a (9-5)
; for h <= a (9-5)
 = experimental semivariogram value at h,
= experimental semivariogram value at h,
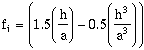 (9-6)
(9-6)
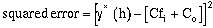 (9-7)
(9-7)
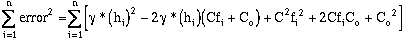 (9-8)
(9-8)
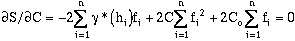
 (9-10)
(9-10)
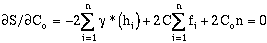
 (9-11)
(9-11)
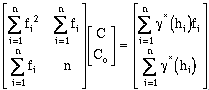 (9-12)
(9-12)
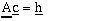

 (9-13a)
(9-13a) (9-13b)
(9-13b) (9-13c)
(9-13c) (9-13d)
(9-13d) (9-13e)
(9-13e)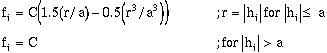 (9-13f)
(9-13f) (9-13g)
(9-13g)  (9-14)
(9-14)
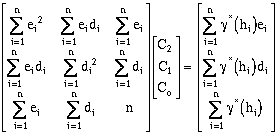 (9-15)
(9-15)
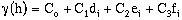 (9-16)
(9-16)
 (9-17)
(9-17)
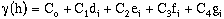 (9-18)
(9-18)
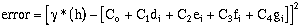 (9-19)
(9-19)
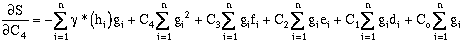


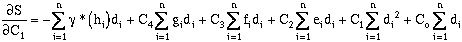
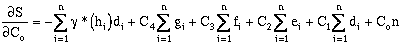

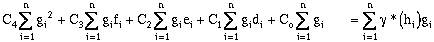 (9-20)
(9-20)
 (9-21)
(9-21)
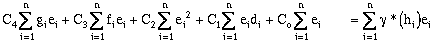 (9-22)
(9-22)
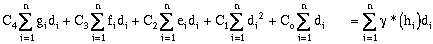 (9-23)
(9-23)
 (9-24)
(9-24)
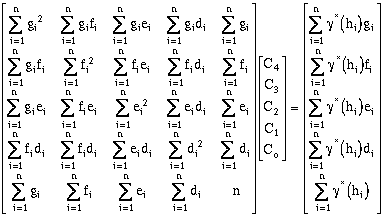 (9-25)
(9-25)
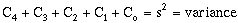 (9-26)
(9-26)
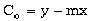 (9-27)
(9-27)
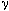 (h) coordinates of the first lag point, and m is the
slope of the line between the shortest two lag points. This isn't necessarily a good
assumption, but on a practical basis, it can improve the model solution in some cases.
Another alternative is to weight each lag point by the number of data pairs that went
into defining that point, the theory being that a point represented by many data points
should be honored more closely then a lag point only based on a few sample pairs.
(h) coordinates of the first lag point, and m is the
slope of the line between the shortest two lag points. This isn't necessarily a good
assumption, but on a practical basis, it can improve the model solution in some cases.
Another alternative is to weight each lag point by the number of data pairs that went
into defining that point, the theory being that a point represented by many data points
should be honored more closely then a lag point only based on a few sample pairs.