

The Grid module interpolates parameter values at locations were there are no physical data. This is done using various interpolation algorithms (inverse-distance, kriging, trend-surface analysis) based on irregularly spaced data. Sometimes it is of interest to estimate what is occurring between data locations. For other applications, for convenience, or for clarity, irregularly spaced data must be interpolated onto a regular grid to be useful. For example contour, surface, and block require that the data being viewed be gridded with a rectangular pattern. These programs then allow the user to visually view the interpolated estimate of the field data. Grid is used to interpolate values at locations of convenient based on field data.
Within grid there are several gridding algorithms; inverse-distance, simple and ordinary kriging method, and trend-surface analysis. Inverse-distance is a relatively simple method which estimates the value of a location based on the distance and value of surrounding points. Kriging does much the same thing as inverse-distance, except kriging also considers spatial statistics describing how the field data varies directionally. Kriging is often referred to a the best unbiased estimator to estimate a value for a given location. Trend-surface analysis is basically a least-squares regression technique which assumes a data value is a function of a "regional" trend and minor "local" variations. The calculated trend-surface attempts to model the "regional" component.
The grid application is composed of two sections (Figure 10.1); the main menu-bar, and the log/status area. The menu-bar is used to select all grid commands and the log/status area contains relevant data about the status of the program or the state of on-going calculations.
The parameters defining the search and weighting parameters are the search radius, tolerance, power, nearest points, search type, and the grid file name. The search radius specifies the maximum a data point can be from the point being gridded and still have an influence on its estimated value; the default value is the maximum diagonal distance across the data set (i.e. all points may influence any grid location). The tolerance is used to handle a problem with the inverse-distance algorithm. If the point being gridded is at the same location as a data point, the distance between the two locations is zero; if this is not specially treated, a division by zero error occurs. The tolerance is used to avoid this problem (See Mathematics Section for further details). Power is a weighting function for the distance between the gridded location and the surrounding data points. If power equals 1.0, the inverse-distance algorithm regresses to a normal linear interpolation. Power set equal to 2.0 is a commonly used weight (See Mathematics Section for details). The nearest points refer to the maximum number of "nearby" points that will be used to estimate a grid location, 10 is a commonly used number. Depending on the data set more or fewer points may be called for.
Currently, the software only supports a normal search; not quadrant or octant searches. In some data sets it is possible to get artificial benching on slopes, and peaks and valleys or pits tend to become flat; these different search methods can reduce these problems (See Mathematics Section).
The grid file is the file where the calculated results will be saved. The file will be formatted for contour (Chapter 11), surface (Chapter 12), and block (Chapter 13) using a GRID CENTERED GRID format (Figure 10.3, see Setting up a "Standard" Input Grid File section (Chapter 11)). The filename can be typed directly in the text field or, the Select button next to the text entry field maybe used. It generates a pop-up dialog similar to that in Figure 5.2. The default search extension for 2D solutions, though is *.srf. The filename may also be selected using the File:Save as menu options (This option, however will also save the file; not just rename the destination). After the grid is calculated, the results can be saved to the grid file by pressing the Save button or by selecting the File:Save menu-bar option.
To calculate the inverse-distance grid, press 1) the Calculate button at the bottom of this pop-up dialog, or 2) press the Calculate button at the bottom of either the bottom of the 2D or 3D pop-up dialogs, or 3) select either the Method:Calculate or the Grid:Calculate menu-bar options. When the calculation is started, the pop-up dialog shown in Figure 10.4 will be displayed. This dialog will be displayed as long as the computer is calculating the grid. When the computation is complete it will be removed. If you want to abort the calculation, press the "Stop" button. As soon as the current column calculation is complete (This may take several second on large grids or large data sets), the calculation will be stopped. Note; if the calculation is aborted, no usable information is retained. While the grid is being calculated, the current column being calculated will be displayed in the log/status window, and "Grid Complete" will be printed when the calculation is done.
WARNING: This module should not be used by those unfamiliar with kriging and semivariogram analysis.
For the kriging algorithm there are many variable take must be set by the user, there are some that can ignored for some data sets, and there are some where the default values are acceptable (there may be better values) in most cases. These parameters are described below. For a complete description of these variable it is suggested that the reader refer to Deutsch and Journel (1992).
The first item to decide on is whether Simple or Ordinary Kriging will be used. If Simple kriging is used, you must enter the data sets mean value (This can be determined in histo). The data set mean is assumed, but if a different value is desired, press the Simple Mean button. The pop-up dialog shown in Figure 10.6 can be used set a new value. Note, it is important to check and set these values correctly if "zonal kriging" is used (discussed below), because the mean for the data set may not be relevant for each zone.
The output file names and the Debug Level can also be specified. Be careful not to save results over previously existing files. A 0 Debug Level gives very little information. A 3 Debug Level can give many megabytes of debugging information.
To set the remaining information, there are several buttons for creating input dialogs for defining search, semivariogram, drift, and zonal parameters.
Finally, to calculate the grid, press the Calculate button at the bottom of Figure 10.4 or select the Method:Calculate menu option.
If "zonal kriging" is used, complete semivariogram model definitions must be defined for each zone.
It is also possible to define different semivariogram models for each axis, though the sills in each direction must be identical in all directions at the maximum range. This allows up to three semivariogram models to be defined for each zone (Principle (X), Y-Axis, and Z-Axis). This takes more work to model, but in some cases, modeling the data, it appears that different directions fit different models (e.g. horizontal is a two- nested spherical, but vertical is exponential). This option can be selected by turning the Semivariogram Anisotropies Off. The normal and default position is On.
If "zonal kriging" is used, the Search Radius, Search Angles (1,2,3) and the Anisotropies must be defined for each zone.
Figure 10.13a
and
Figure 10.13b
The Model Semivariograms button creates the same pop-up dialog as shown in Figure 10.7 and described above. The Select Zone Definition Files pop-up dialog is shown in Figure 10.14. This file is used to select and view the mask files for each zone (The file format is discussed in the Setting up the Input File section below).
The Relationship Between Zones button creates the dialog shown in Figure 10.15. This dialog allows the user to specify how data points in one zone will be used, will be used when they are in the search neighborhood of a point being kriged in another zone. A Sharp transition means that only points from the specified zone will be ignored (Figure 10.16a). A Gradational transition means that points will be shared across the specified zones (Figure 10.16b). Note, at the transition from one zone to another, the transition can still be abrupt. This can happen in neighboring cells because the model semivariograms are different and the search neighborhood, and there fore point used in the estimation, can be substantially different. This problem will tend to disappear with increasing model ranges and sample data densities. The Fuzzy transition is not yet supported. Note that this relationship matrix can be loaded or saved using the dialog in Figure 10.12 (The file format is discussed in the Setting up the Input File section below).
Figure 10.16a
and
Figure 10.16b
For the trend-surface algorithm there is only one parameter, the surface order. The remaining variables, ANOVA File, Grid File, and Residual File, simply define where the results will be saved. Valid entries for Order are one to five; higher order surfaces are not supported; the solution matrix size becomes prohibitive.
data points in the data set. This is the number of unknowns in the solution equation.
Filenames for the ANOVA, Grid, or Residual files may be entered in the appropriate text field, or by pressing the appropriate Select button a pop-up dialog (similar to Figure 5.2) will be created allowing the output file to be selected. The default data file extensions are:
To save the data after a calculation is made, press each of the appropriate Save buttons.
The ANOVA file will store statistical information about the trend-surface (the same data printed to the log-status window). The Grid file will store the gridded "regional" trend- surface. The Residual file will store six columns of data; the X, Y, Z, location and observed value for each data point, the residual (actual - estimate), and the trend - surface estimate at each data location.
To calculate the trend-surface grid, press 1) the Calculate button at the bottom of this pop-up dialog, or 2) press the Calculate button at the bottom of either the bottom of the 2D or 3D pop-up dialogs, or 3) select either the Method:Calculate or the Grid:Calculate menu-bar options. When the calculation is started, the pop-up dialog shown in Figure 10.4 will be displayed. This dialog will be displayed as long as the computer is calculating the grid (This is a fairly fast calculation). When the computation is complete "Trend-Surface Grid Complete" will be printed in the log-status window.
When the trend-surface is calculated the trend-surface parameters and several statistical parameters are printed to the log-status window. The statistical variables are calculated are used to evaluate the "goodness of fit" of the calculated surface to the field data. These variables are summarize here, but further details may be found in the Mathematics section of this chapter, or in Davis (1986). The statistical analysis is referred to as the Analysis of Variance (ANOVA). The terms of interest are:
The two main variables of concern are the R2 and the F-statistic. The R2 term when multiplied by 100% defines the amount of data variation explain by the regression trend-surface (Davis, 1986).
The Fstatistic is used to evaluate whether all of the coefficients as a group are significantly different then zero, and the surface is considered meaningful. All of these terms and their use are defined and explained in the Mathematics section. Suffice it to say here though that the best trend-surface will maximize R2 and minimize the F-statistic.
By default, the program assumes the X data is in column 1, the Y data in column 2, and the Z (elevation) data is in column 3. If your data file is prepared differently, or you have multiple Z data columns you will have to redefine the column definitions.
The Grid Dimensions area is where the density and the extent of the grid is specified. The number of Rows and Columns desired is of user preference. As a rule of thumb though a denser grid (more rows and columns) will generate a smoother, but not necessarily a more accurate surface. The X and Y Minimum and Maximum values are by default set the extents of the X and Y data. Depending on the area of interest within the gridded data, or simply shifting these values to whole numbers (e.g. 0.19 is close to 0.0) so the coordinate labels on contour maps are not to busy, the grid extents are user definable.
From the menu bar select File:Open, and select the file "conc3.dat." This opens the data file. By viewing the data file (File:View:Data), it can be seen that this a 2D data set even though X, Y, Z, and Value data are supplied. The Y data column (column 2) contains only zeros. To grid this data, the 2D gridding option will be used where column 1 (X data referring to horizontal distance), column 3 (Z data referring to elevation above some datum), and column 4 (Value data representing contaminant concentrations in ppm) are used. Use File:Quit in the editor to destroy the editor window.
To set up the gridding calculation, select from the menu-bar, Method:Inverse- Distance. This accomplishes two tasks; 1) it tells the program to use the Inverse- Distance solver when the grid is calculated, and 2) it creates the pop-up dialog shown in Figure 10.2 allowing parameters about the gridding method to be defined. For this example the default Solution Parameters and the Search Type are reasonable.
Next, select the Grid:2D Grid menu-bar option. The pop-up dialog shown in Figure 10.18 will be created. The Y Column, however needs to be redefined. Since this data set is a cross-section (X-Z plane) we need to map the Z data to the Y-axis on the grid (the grid always grids using the X and Y axes). In this case the Y Column should specify the Z data column in the file; enter 3 in the Y Column field. The Value Column now specifies data column 3; this needs to be changed to column 4. When you hit <RETURN> in each field, or hit the Apply dialog button, the Minimum and Maximum File Data values should appear as:
X Column: 0.00 to 350.0 Y Column: 12.5 to 139.0 Value Column: -0.0175 to 100.0
The Grid Dimensions are reasonable, but if you wanted the grid cells to be approximately square, you could change the number of Columns (X) to 50 (grid cell 7.14' x 5.23'). Exit the dialog by pressing the Done dialog button.
To calculate the grid, select Grid:Calculate from the menu-bar, or press either Calculate button on the Inverse-Distance dialog or the 2D Grid dialog. This starts the calculation process. In the text area of the main window the status of the solution is displayed. As each column is calculated it is printed. When the calculation is completed, the message "Grid Complete" is displayed. Note that the program has only calculated the grid; no results have been saved to a file! To save the results, use the File:Save or File:Save as menu-bar options, or the Save button in the Inverse-Distance dialog.
To view the results, select form the menu-bar View:Contour Map or View:Surface Map. If the results have been saved, contour (Chapter 11) or surface (Chapter 12) is called and the gridded results of the data set are mapped. If the results haven't been saved you will be requested to save the results. If no file has been previously saved, you will be queried for a file name, otherwise the results will be saved to the file used last (i.e. the previous results will be overwritten).
The default 2D grid parameters will be reasonable. Next, open the Method:Trend- Surface Analysis pop-up dialog. The dialog should come up with a first-order trend (a plane) specified. Press the Calculate button. In the log-status window the regression equation and the ANOVA results will be printed. The trend-surface equation for any X- Y point is:
The squared Multiple Correlation Coefficient (R2), in this example suggest that the trend-surface explains 30.06% of the data set variation. The Fstatistic of 93.83 with 3/655 Degrees of Freedom , using an F-Test with a 1% (a = 0.01) Level of Significance, implies that the NULL hypothesis (H0, the terms of the regression are not significantly different then zero) is not true (93.83 > 3.78, Table 10.1c), therefore this regression has value in describing the nature of the surface. Next, save the regression surface (press the Grid File Save button). Once the file is saved, to make a contour map of the first order trend, select the menu-bar option View:Contour Map. This will call the program contour (Chapter 11) and show the "regional" trend-surface (Figure 10.23a).
To examine the "local" variations between the observed data and the "regional" trend surface, save the Residual File. Next select File:Open, and open the file you just saved (default name is junk.res.dat). The data residuals are stored in column 5, so bring up the 2D Grid dialog (Grid:2D Grid) and change the Value Column from 4 to 5. This data needs to be gridded using conventional gridding algorithm. For this example use inverse-distance. Select the Method:Inverse-Distance menu-bar option. The default values are reasonable in the new dialog. Press Calculate button in the Inverse-Distance dialog. Once it is done calculating, press the Save button for the inverse-distance Grid File. Selecting the View:Contour Map will start another version of contour with the residuals mapped (Figure 10.23b).
This process can be repeated for each trend-surface order, two through five. The resultant trend-surface and residual maps are shown in Figures 24a & 24b, Figures 25a & 25b, Figures 26a & 26b, and Figures 27a & 27b. The results are summarized below.
Figure 10.24a and
Figure 10.24b
Figure 10.25a and
Figure 10.25b
Figure 10.26a and
Figure 10.26b
Figure 10.27a and
Figure 10.27b
2nd order:
Regression explains 44.95% of variation (R2 = 0.4495)
3rd order:
Total F-Test = 148.3 D.F. = ( 10 / 648 )
Regression explains 69.59% of variation (R2 = 0.6959)
4th order:
Total F-Test = 136.4 D.F. = ( 15 / 643 )
Regression explains 76.09% of variation (R2 = 0.7609)
5th order:
Total F-Test = 153.2 D.F. = ( 21 / 637 )
Regression explains 83.47% of variation (R2 = 0.8347)
For each of these regressions, the Fstatistic with a 1% (a = 0.01) Level of Significance, implies that the NULL hypothesis is not true, therefore this regression has value in describing the nature of the surface.
Syntax:
Meaning of flag symbols:
NOTES:
If no entry is required for flag, flag command executed.
Flag Definitions:
| -anovae | = | ANOVA file search extension | default = "*.anova" | ||
| -anovaf | = | ANOVA file name | default = "junk.anova" | ||
| -cl | = | number of grid columns (Y) | default = 25 | ||
| -dft | = | drift option | default = 0 | ||
|
|||||
| -dfte | = | drift search extension | default = "*.dft" | ||
| -dftf | = | drift file | default = "junk.dft" | ||
| -dfti | = | drift input column | default = 5 | ||
| -dfto | = | drift output column | default = 4 | ||
| -do {} | = | drift output column | default = 0 | ||
|
|||||
| -dfte | = | estimation search extension | default = "*.est" | ||
| -dftf | = | estimation file | default = "junk.est" | ||
| -gd | = | grid dimension | default = 0 | ||
|
|||||
| -help | = | give this help menu | |||
| -kdbge | = | kriging debug search extension | default = "*.dbg" | ||
| -kdbgf | = | kriging debug file | default = "junk.dbg" | ||
| -kdbgl | = | kriging debug level | default = 0 | ||
| -kst | = | kriging search method | default = 0 | ||
|
|||||
| -kt | = | kriging method | default = 1 | ||
|
|||||
| -lgf | = | log file name | defalut = "log.dat" | ||
| -ly | = | number of grid layers (Z) | default = 1 | ||
| -m1 {} | = | Model types nest #1 | default = 3 | ||
|
|||||
| -m2 {} | = | model types nest #2 (See -ml) | default = 0 | ||
| -m3 {} | = | model types nest #3 (See -ml) | default = 0 | ||
| -m4 {} | = | model types nest #4 (See -ml) | default = 0 | ||
| -mode | = | semivariogram model search extension | default = "*.mod" | ||
| -modf {} | = | semivariogram model file | default = " " | ||
| -ng {} | = | nugget | default = 0.0 | ||
| -nm | = | number of semivariogram models | default = 1 | ||
| -np | = | number of nearest search neighbors | default = 10 | ||
| -npmax | = | maximum number of points | default = 16 | ||
| -npmin | = | minimum number of points | default = 4 | ||
| -nz | = | number of zones | default = 1 | ||
| -octmax | = | maximum points per octant | default = 4 | ||
| -out | = | output *.out filename | defalut = " " | ||
| -pr | = | inverse distance power | default = 2.0 | ||
| -prf | = | preference file name | defalut = "grid.prf" | ||
| -r1 {} | = | range for nest #1 | default = 0.0 | ||
| -r2 {} | = | range for nest #2 | default = 0.0 | ||
| -r3 {} | = | range for nest #3 | default = 0.0 | ||
| -r4 {} | = | range for nest #4 | default = 0.0 | ||
| -s1 {} | = | C for nest #1 | default = 0.0 | ||
| -s2 {} | = | C for nest #2 | default = 0.0 | ||
| -s3 {} | = | C for nest #3 | default = 0.0 | ||
| -s4 {} | = | C for nest #4 | default = 0.0 | ||
| -rese | = | residual file search extension | default = "*.res.dat" | ||
| -resf | = | residual file name | default = "junk.res.dat" | ||
| -run | = | run and calcualte grid without X-interface | |||
| -rw | = | number of grid rows (X) | default = 25 | ||
| -sang1 {} | = | neighbor search angle 1 | default = 0.0 | ||
| -sang2 {} | = | neighbor search angle 2 | default = 0.0 | ||
| -sang3 {} | = | neighbor search angle 3 | default = 0.0 | ||
| -sn | = | kriging method | default = 0 | ||
|
|||||
| -so | = | trend-surface order number | default = 1 | ||
| -sr | = | searc h radius | default = max. diagonal | ||
| -st | = | solution type | default = 0 | ||
|
|||||
| -syaniso {} | = | neighbor anisotropy 1 | default = 1.0 | ||
| -szaniso {} | = | neighbor anisotropy 2 | default = 1.0 | ||
| -tl | = | tolerence | default = 1/10 grid spacing | ||
| -tmmax | = | maximum trimming limit | default = 1.0e30 | ||
| -tmmin | = | minimum trimming limit | default = -1.0e30 | ||
| -va11 {} | = | angle 1 for nest #1 | default = 0.0 | ||
| -va12 {} | = | angle 1 for nest #2 | default = 0.0 | ||
| -va13 {} | = | angle 1 for nest #3 | default = 0.0 | ||
| -va14 {} | = | angle 1 for nest #4 | default = 0.0 | ||
| -va21 {} | = | angle 2 for nest #1 | default = 0.0 | ||
| -va22 {} | = | angle 2 for nest #2 | default = 0.0 | ||
| -va23 {} | = | angle 2 for nest #3 | default = 0.0 | ||
| -va24 {} | = | angle 2 for nest #4 | default = 0.0 | ||
| -va31 {} | = | angle 3 for nest #1 | default = 0.0 | ||
| -va32 {} | = | angle 3 for nest #2 | default = 0.0 | ||
| -va33 {} | = | angle 3 for nest #3 | default = 0.0 | ||
| -va34 {} | = | angle 3 for nest #4 | default = 0.0 | ||
| -vc | = | value data column | default = 4 | ||
| -xbdes | = | x block descretization | default = 1 | ||
| -xc | = | x data column | default = 1 | ||
| -xmax | = | maximum gridded X value | default = maximun data X | ||
| -xmin | = | minimum gridded X value | default = minimun data X | ||
| -ya1 {} | = | anisotropy 1 for nest #1 | default = 1.0 | ||
| -ya2 {} | = | anisotropy 1 for nest #2 | default = 1.0 | ||
| -ya3 {} | = | anisotropy 1 for nest #3 | default = 1.0 | ||
| -ya4 {} | = | anisotropy 1 for nest #4 | default = 1.0 | ||
| -ybdes | = | y block descretization | default = 1 | ||
| -yc | = | y data column | default = 2 | ||
| -ymax | = | maximum gridded Y value | default = maximun data Y | ||
| -ymin | = | minimum gridded Y value | default = minimun data Y | ||
| -za1 {} | = | anisotropy 2 for nest #1 | default = 1.0 | ||
| -za2 {} | = | anisotropy 2 for nest #2 | default = 1.0 | ||
| -za3 {} | = | anisotropy 2 for nest #3 | default = 1.0 | ||
| -za4 {} | = | anisotropy 2 for nest #4 | default = 1.0 | ||
| -ybdes | = | z block descretization | default = 1 | ||
| -zc | = | z data column | default = 3 | ||
| -zc | = | zone search extension | default = "*.zon" | ||
| -zf | = | zone file | default = " " | ||
| -zmax | = | maximum gridded Z value | default = maximun data Z | ||
| -zmin | = | minimum gridded Z value | default = minimun data Z | ||
| -zrc | = | zone relationship search extension | default = "*.zrl" | ||
| -zrf | = | zone relationship file | default = " " |
For an example, see data file grad.zrl.
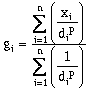 (10-1)
(10-1)where gi is the estimated value at the grid location, di is the distance between the grid location and the sample data, and p is the power to which the distance is raised. The basis of this technique is that nearby data are most similar to the actual field conditions at the grid location. Depending on the site conditions the distance may be weighted in different ways. If p = 1, this is a simple linear interpolation between points. Many people have found that p = 2 produces better results. In this case, close points are heavily weighted, and more distant point are lightly weighted (points are weighted by 1 / d2). At other sites, p has been set to other powers and yielded reasonable results.
Inverse-distance is a simple and effective way to estimate parameter values at grid locations of unknown value. Beside the directness and simplicity, however, inverse-distance techniques have a number of shortcomings. Some of these are:
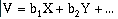
where V is the data value at the described location, the b's are coefficients, and X and Y are combinations of geographic location.
An example is shown in Figure 10.28. For a set of data, a line or surface may be fit exactly through each point, but a trend (a mathematical equation) can be defined which approximates the data well "regionally" with only minor "local" variations.
Trend-surface analysis is basically a linear regression technique, but it is applied to two- and three-dimensions instead of just fitting a line. A first order linear trend surface equation has the form:
 (10-2)
(10-2)That is, an observation, with a value V, can be described as a linear function of a constant value (b0) related to the data set mean, and east-west (b1), and north-south (b2) components (Davis, 1973). To solve for these three unknowns, three normal equations are available (Davis, 1973):
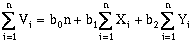 (10-3)
(10-3)
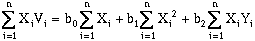 (10-4)
(10-4)
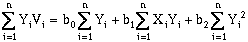 (10-5)
(10-5)
where n is the number of data points. Solving these equations simultaneously will yield a "best-fit", defined by least-squares regression, for a two-dimensional, first-order (a plane) trend surface. This can be rewritten in matrix format:
 (10-6)
(10-6)This approach can also be applied to higher order surfaces and three-dimensions. In three-dimensions, the equivalent equation would be:
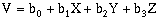 (10-7)
(10-7)
 (10-8)
(10-8)
For second-order, two-dimensional surfaces, the general equation is:
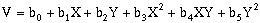 (10-9)
(10-9)Third-order, two-dimensional surfaces:
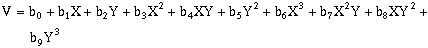 (10-10)
(10-10)Forth-order, two-dimensional surfaces:
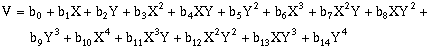 (10-11)
(10-11)Fifth-order, two-dimensional surfaces:
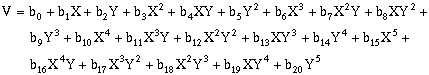 (10-12)
(10-12)For second-order, three-dimensional surfaces, the general equation is:
 (10-13)
(10-13)Third-order, three-dimensional surfaces:
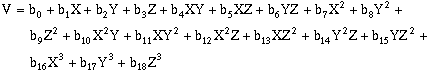 (10-14)
(10-14)Forth-order, three-dimensional surfaces:
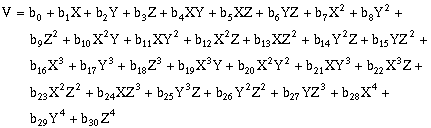 (10-15)
(10-15)Fifth-order, three-dimensional surfaces:
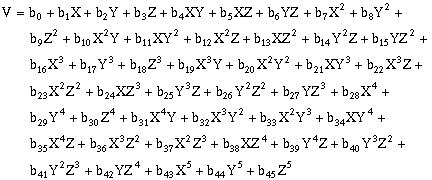 (10-16)
(10-16)
where:
 | (10-18) | |
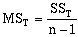 | (10-19)) | |
 | (10-20)) | |
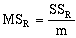 | (10-21)) | |
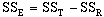 | (10-22)) | |
 | (10-23)) | |
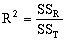 | (10-24) | |
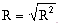 | (10-25) | |
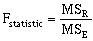 | (10-26) |
where n equals the number of data observations, and m equals the number of coefficients in the trend-surface equation. R, R2 and the Fstatistic are terms used to evaluate the "goodness of fit." R is referred to as the coefficient of multiple correlation and R2 x 100% effects the percent of the data variation explained by the regional trend- surface (Davis, 1986). The Fstatistic is used with the F-test to determine if the group of trend-surface coefficients are significantly different than zero; i.e. the regression effect is not significantly different from the random effect of the data. In formal statistical terms, the F-test for significance of fit tests the hypothesis (H0) and alternative (H1):
 (10-27)
(10-27)The hypothesis tested, is that the partial regression coefficients equal zero, i.e. there is no regression. If the computed F value exceeds the table value of F (Tables 10.1a, 10.1b, and 10.1c), the NULL hypothesis is rejected and the alternative is accepted, i.e. all the coefficients in the regression are significant and the regression is worthwhile.
Table 10.1a,
Table 10.1b, and
Table 10.1c
In addition to the problems of getting a "good" fit for the trend surface, there are a number of pit-falls to the technique (Davis, 1986):
Burrough, P.A., 1986, Principles of Geographical Information Systems for Land Resource Assessment, Monographs on Soil and Resources Survey No. 12, Oxford Science Publications - Clarendon Press, Oxford.
Davis, John C., 1986 (Second Edition), Statistics and Data Analysis in Geology, John Wiley & Sons, New York, pp 405-425.
Deutsch, C.V., and A.G. Journel, 1992, GSLIB: Geostatistical Software Library and User's Guide, Oxford University Press, New York.
Kirk, K.G., 1991, Residual Analysis for Evaluating the Robustness of Inverse Distance, Kriging, and Minimum Tension Gridding Algorithms, GeoTech/GeoChautauqua 91' Conference Proceedings, Lakewood, Colorado, pg. 15 (presentation).
Wingle, W.L., 1992, Examining Common Problems Associated with Various Contouring Methods, Particularly Inverse-Distance Methods, Using Shaded Relief Surfaces, Geotech '92 Conference Proceedings, 1992, Lakewood, Colorado.